Our research
The vulnerability of cancer cells to their addiction to kinase activities
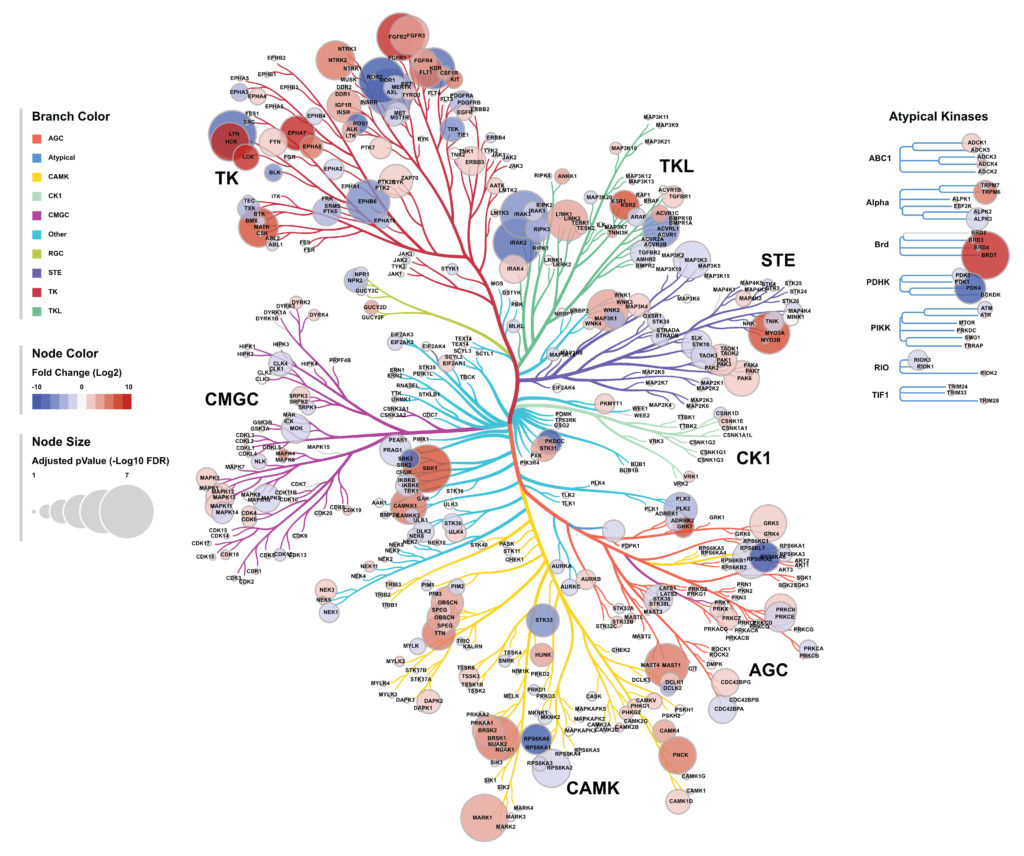 In decades, the survival rate of several cancers has barely been improved. The survival and the proliferation of such cancers strongly rely on a network of kinases of which the activities fluctuate. Every tumors harbor a variety of genetic alterations directly or indirectly affecting the activity of kinases involved in signaling pathways, the cell cycle or the proliferation. However, the heterogeneity of alterations in such tumors has rendered difficult the development of a unique therapeutic strategy to treat a unique cancer type. It is therefore essential to evaluate how tumors are addicted to kinases and their activities parallel to specific oncogenic alterations.
In decades, the survival rate of several cancers has barely been improved. The survival and the proliferation of such cancers strongly rely on a network of kinases of which the activities fluctuate. Every tumors harbor a variety of genetic alterations directly or indirectly affecting the activity of kinases involved in signaling pathways, the cell cycle or the proliferation. However, the heterogeneity of alterations in such tumors has rendered difficult the development of a unique therapeutic strategy to treat a unique cancer type. It is therefore essential to evaluate how tumors are addicted to kinases and their activities parallel to specific oncogenic alterations.
Our research focuses on assessing the addiction of cancer cells to kinase activities and predict their response to kinases inhibitors using a multiomics approach. At this end, our lab takes advantage of various in vitro and in vivo models especially of oesophageal squamous carcinoma to challenge tumor cells to their kinase addiction.
![]()
As the CDK4 activity lies downstream from most of the oncogenic deregulations of signalling pathways, it has been proposed to represent the therapeutic target of choice. Despite this, not all tumors respond to CDK4 inhibition and long-term treatment triggers escape mechanisms. In this context, it is crucial to characterize the sensitivity of tumors including eSCC to CDK4 inhibition, understand the CDK4 inhibition on the cell cycle progression and chromosome instability to fully decipher the molecular response for effective combined targeted therapy.
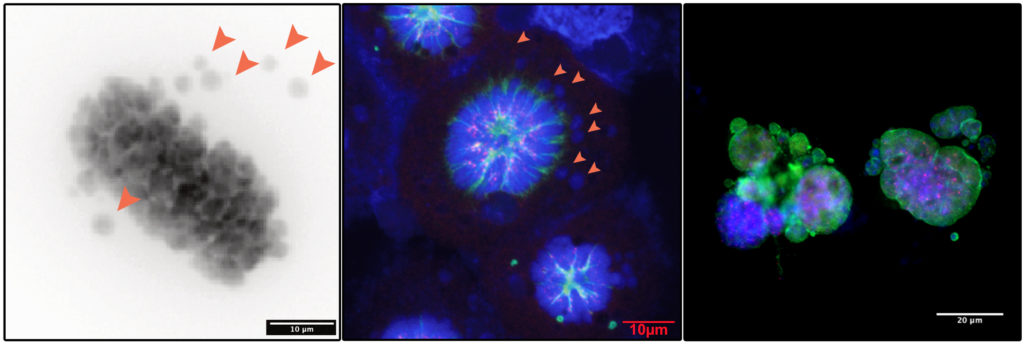
We try to address all these questions using various approaches combining:
- Mass spectrometry based proteomics and phosphoproteomics
- Bidimensional electrophoresis and kinase assays
- NGS and bioinformatics
- Transgenic mouse models
- 2D/3D cell cultures of cancer cell lines/isolated mouse tumor/patient tumor derived cells
- Immunofluorescence and time lapse imaging
Funded by MSCA action and Innoviris Brain 2 Brussels – Attract program
Group members
Group members
Xavier Bisteau, PhD,
Group leader (Xavier.Bisteau@ulb.ac.be)
phone # +32 2 555 4153

Xavier Bisteau graduated as a molecular biologist (MS in 2007) and received his PhD in biomedical and pharmaceutical sciences from the Université Libre de Bruxelles (ULB) in 2013. He performed a postdoctoral training in the lab of Prof. Philipp Kaldis at IMCB (A*STAR) in Singapore from 2013 to 2019. Thanks to 2 fundings (Marie Curie fellowship and Innoviris – Attract), he returned as independent research fellow in 2019. He officially opened his lab in 2021 following his appointment to a tenure position as “chargé de cours” (Associate Professor) in 2021
![]()
Fabiana Moresi, PhD student
(Fabiana.Moresi@ulb.be)

Fabiana obtained a double Master degree in Genetics & Molecular Biology from La Sapienza in Rome and from Paris-Diderot/Paris-Descartes in 2020. She joined the lab in November 2020 to study the interplay between cancer cells, the immune system and the genomic instability following CDK4/6 inhibition
![]()
Marta Avalo, PhD student
(marta.avalos.morenoi@ulb.be)
 Marta received her master degree in 2020 from the University of Granada. She further studied bioinformatics at the KU Leuven before joining the lab in November 2021. Marta is studying the dynamic changes of the protein-protein interaction network parallel to phosphorylation modulations following kinase inhibition in cancer.
Marta received her master degree in 2020 from the University of Granada. She further studied bioinformatics at the KU Leuven before joining the lab in November 2021. Marta is studying the dynamic changes of the protein-protein interaction network parallel to phosphorylation modulations following kinase inhibition in cancer.
![]()
Virginie Imbault, MS
Virginie received her master in genetics and microbiology in 2004 and a professional master in proteomics in 2005 from the University of Lille 1. She joined the institute in 2006 and is the technical manager of the proteomics facility. She is working in collaboration with our group, the group of David Communi and all collaborators interested in MS-based proteomics analyses.
Publications
22 Redondo Marin, J. A., Bibes, R., Vercauteren Drubbel, A., Dassy, B., Bisteau, X., Maury, E., & Beck, B. (2021). PER2 Circadian Oscillation Sensitizes Esophageal Cancer Cells to Chemotherapy. Biology, 10(4), 266. doi:10.3390/biology10040266
21 Dewhurst MR, Ow JR, Zafer G, van Hul NKM, Wollmann H, Bisteau X, Brough D, Choi H, Kaldis P. (2020) Loss of hepatocyte cell division leads to liver inflammation and fibrosis. PLoS Genet. 16(11):e1009084. doi: 10.1371/journal.pgen.1009084.
20 Bisteau X*, Lee J, Srinivas V, Lee JHS, Niska-Blakie J, Tan G, Yap SYX, Hom KW, Wong CK, Chae J, Wang LC, Kim J, Rancati G, Sobota RM, Tan CSH, Kaldis P. (2020) The Greatwall kinase safeguards the genome integrity by affecting the kinome activity in mitosis. Oncogene. (44):6816-6840. doi: 10.1038/s41388-020-01470-1 *corresponding author
19 Palmer N, Talib SZA, Ratnacaram CK, Low D, Bisteau X, Lee JHS, Pfeiffenberger E, Wollmann H, Tan JHL, Wee S, Sobota R, Gunaratne J, Messerschmidt DM, Guccione E, Kaldis P. (2019) CDK2 regulates the NRF1/Ehmt1 axis during meiotic prophase I. J Cell Biol. 218(9):2896-2918. doi: 10.1083/jcb.201903125.
18 Szmyd R., Niska-Blakie J., Diril M.K., Renck Nunes P., Tzelepis K., Lacroix A., van Hul N., Deng LW., Matos J., Dreesen O., Bisteau X., Kaldis P. (2018) Premature activation of Cdk1 leads to mitotic events in S phase and embryonic lethality Oncogene. 2019 Feb;38(7):998-1018. doi: 10.1038/s41388-018-0464-0
17 Dai L., Zhao T., Bisteau X., Sun W., Prabhu N., Lim YT., Sobota R., Kaldis P., Nordlund P. (2018) Modulation of protein interaction states through the cell cycle. Cell 173(6):1481-1494.e13 doi: 10.1016/j.cell.2018.03.065
16 Tan CSH., Go KD., Bisteau X., Dai L., Yong CH., Prabhu N., Ozturk MB., Lim YT., Sreekumar L., Lengqvist J., Tergaonkar V., Kaldis P., Sobota S., Nordlund P. (2018) Thermal Proximity Co-aggregation for System-wide Monitoring of Protein Complex Dynamics in Intact Cell. Science 359(6380):1170-1177 doi: 10.1126/science.aan0346
15 Windpassinger, C., Piard, J., Bonnard, C., Alfadhel, M., Lim, S., Bisteau, X., et al. (2017). CDK10 Mutations in Humans and Mice Cause Severe Growth Retardation, Spine Malformations, and Developmental Delays. American Journal of Human Genetics, 101(3), 391–403
14 Colleoni, B., Paternot, S., Pita, J. M., Bisteau, X., Coulonval, K., Davis, R. J., et al. (2017). JNKs function as CDK4-activating kinases by phosphorylating CDK4 and p21. Oncogene, 9, 391. http://doi.org/10.1038/onc.2017.7
13 Kim, S. Y., Lee, J.-H., Merrins, M. J., Gavrilova, O., Bisteau, X., Kaldis, P., et al. (2017). Loss of Cyclin-dependent Kinase 2 in the Pancreas Links Primary β-Cell Dysfunction to Progressive Depletion of β-Cell Mass and Diabetes. The Journal of Biological Chemistry, 292(9), 3841–3853. http://doi.org/10.1074/jbc.M116.754077
12 Jayapal, S. R., Ang, H. Y.-K., Wang, C. Q., Bisteau, X., Caldez, M. J., Xuan, G. X., et al. (2016). Cyclin A2 regulates erythrocyte morphology and numbers. Cell Cycle (Georgetown, Tex.), 1–12. http://doi.org/10.1080/15384101.2016.1234546
11 Chauhan, S., Diril, M. K., Lee, J. H. S., Bisteau, X., Manoharan, V., Adhikari, D., et al. (2016). Cdk2 catalytic activity is essential for meiotic cell division in vivo. The Biochemical Journal, 473(18), 2783–2798. http://doi.org/10.1042/BCJ20160607
10 Diril, M. K*., Bisteau, X.*, Kitagawa, M., Caldez, M. J., Wee, S., Gunaratne, J., et al. (2016). Loss of the Greatwall Kinase Weakens the Spindle Assembly Checkpoint. PLoS Genetics, 12(9), e1006310. http://doi.org/10.1371/journal.pgen.1006310.s015 * Co-first author
9 Heijink, A. M., Blomen, V. A., Bisteau, X., Degener, F., Matsushita, F. Y., Kaldis, P., et al. (2015). A haploid genetic screen identifies the G1/Sregulatory machinery as a determinant ofWee1 inhibitor sensitivity. Proceedings of the National Academy of Sciences of the United States of America, 201505283. http://doi.org/10.1073/pnas.1505283112
8 Jayapal, S. R., Wang, C. Q., Bisteau, X., Caldez, M. J., Lim, S., Tergaonkar, V., et al. (2015). Hematopoiesis specific loss of Cdk2 and Cdk4 results in increased erythrocyte size and delayed platelet recovery following stress. Haematologica, 100(4), 431–438. http://doi.org/10.3324/haematol.2014.106468
7 Bisteau, X., & Kaldis, P. (2014). Spy1/SpeedyA accelerates neuroblastoma. Oncotarget, 5(16), 6554–6555.
6 Bisteau, X., Caldez, M. J., & Kaldis, P. (2014). The Complex Relationship between Liver Cancer and the Cell Cycle: A Story of Multiple Regulations. Cancers, 6(1), 79–111. http://doi.org/10.3390/cancers6010079
5 Paternot, S., Colleoni, B., Bisteau, X., & Roger, P. P. (2014). The CDK4/CDK6 inhibitor PD0332991 paradoxically stabilizes activated cyclin D3-CDK4/6 complexes. Cell Cycle (Georgetown, Tex.). http://doi.org/10.4161/15384101.2014.946841
4 Farhang Ghahremani, M., Goossens, S., Nittner, D., Bisteau, X., Bartunkova, S., Zwolinska, A., et al. (2013). p53 promotes VEGF expression and angiogenesis in the absence of an intact p21-Rb pathway. Cell Death and Differentiation, 20(7), 888–897. http://doi.org/10.1038/cdd.2013.12
3 Bisteau, X.*, Paternot, S.*, Colleoni, B., Ecker, K., Coulonval, K., De Groote, P., et al. (2013). CDK4 T172 phosphorylation is central in a CDK7-dependent bidirectional CDK4/CDK2 interplay mediated by p21 phosphorylation at the restriction point. PLoS Genetics, 9(5), e1003546. http://doi.org/10.1371/journal.pgen.1003546.s011 * equal contribution
2 Paternot, S., Bockstaele, L., Bisteau, X., Kooken, H., Coulonval, K., & Roger, P. P. (2010). Rb inactivation in cell cycle and cancer: the puzzle of highly regulated activating phosphorylation of CDK4 versus constitutively active CDK-activating kinase. Cell Cycle (Georgetown, Tex.), 9(4), 689–699.
1 Bockstaele, L*., Bisteau, X.*, Paternot, S., & Roger, P. P. (2009). Differential regulation of cyclin-dependent kinase 4 (CDK4) and CDK6, evidence that CDK4 might not be activated by CDK7, and design of a CDK6 activating mutation. Molecular and Cellular Biology, 29(15), 4188–4200. http://doi.org/10.1128/MCB.01823-08 * Co-first author